# TruSeq Small RNA v1.0
## Overview
TruSeq Small RNA v1.0 includes the following functionality:
* Preconfigured TruSeq Small RNA v1.0 protocol that explains how to prepare total RNA or purified small RNA using a Illumina® TruSeq® Small RNA Library Prep Kit.
* Automated calculation of sample and buffer volumes.
* Automated calculation or display of reagents at every step in the protocol.
* Automatic step transition when required.
* Automatic placement of samples when necessary.
* Automated assignment of QC Pass/Fail, based on user-selected threshold values.
* A routing script that allows sequencing of libraries using any Illumina sequencing instrument.
It is not required to have the total RNA samples quantified prior to starting this protocol, however it is highly recommended to ensure the starting number of samples.
## Protocol 1: TruSeq Small RNA v1.0
Protocol Type = Library Prep
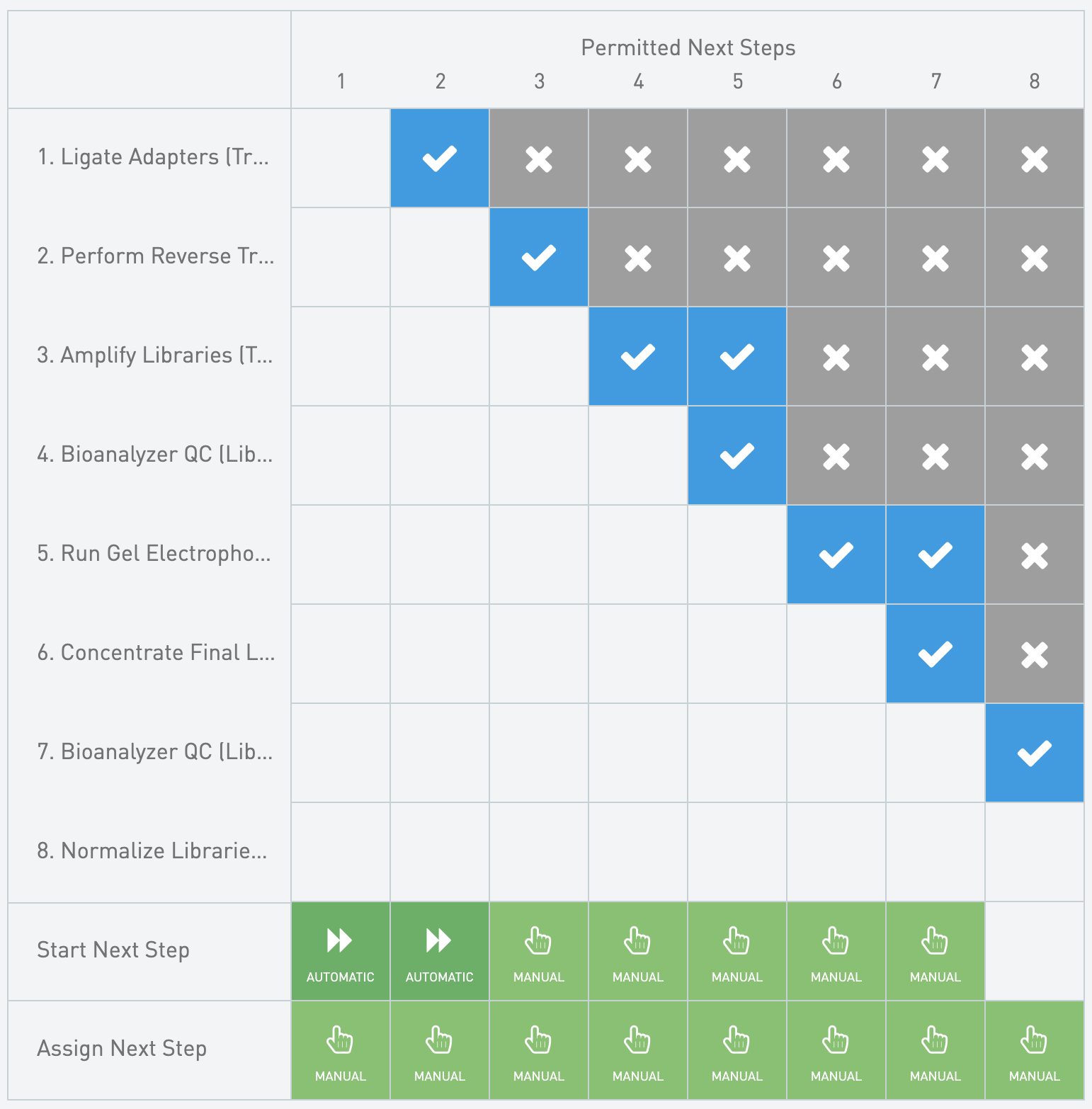
**Next Steps Configuration**
* TruSeq Small RNA Library Prep Kit, Indices A, B, C or D
* Supplier = Illumina
* Catalog Number = RS-200-0012; RS-200-0024; RS-200-0036; RS-200-0048
* Website =
* Control Types
* Human Brain Total RNA
* Supplier = Thermo Fisher Scientific
* Catalog Number = AM7962
{% hint style="info" %}
The version of Ligate Adapters master step name may be different depending on the version of IPP installed.
{% endhint %}
#### Automations
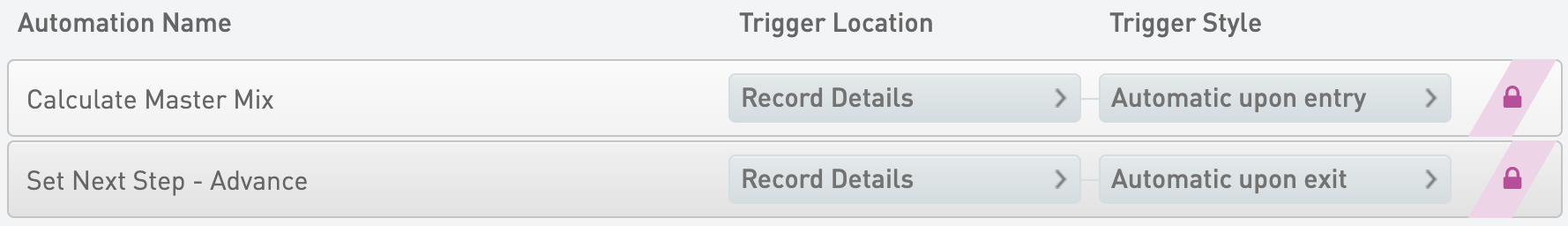
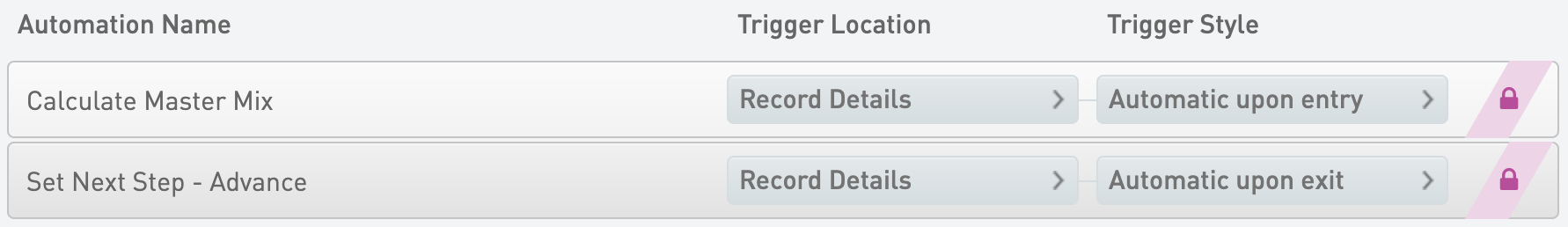
Calculate Master Mix
* Trigger Location = Record Details
* Trigger Style = Automatic upon entry
{% code overflow="wrap" %}
```markup
bash -l -c " /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t false -h false -exp '(step.::Total samples:: = step.::Total samples:: + 1)' -log {compoundOutputFileLuid0} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t false -h false -exp '(step.::HML Volume (ul):: = step.::Total samples:: * 2.2) ; (step.::RNase Inhibitor Volume (ul):: = step.::Total samples:: * 1.1) ; (step.::T4 RNA Ligase 2, Depletion Mutant Volume (ul):: = step.::Total samples:: * 1.1) ; (step.::RA5 Volume (ul):: = step.::Total samples:: * 1.21) ; (step.::10mM ATP Volume (ul):: = step.::Total samples:: * 1.21) ; (step.::T4 RNA Ligase Volume (ul):: = step.::Total samples:: * 1.21)' -log {compoundOutputFileLuid1}"
```
{% endcode %}
Set Next Step - Advance
* Trigger Location = Record Details
* Trigger Style = Automatic upon exit
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} \
script:evaluateDynamicExpression \
-t false \
-h false \
-exp 'nextStep = ::ADVANCE::' \
-log {compoundOutputFileLuid0}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------ | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | LIMS ID (Container) | Built-in | | |
| Project | Project Name | Built-in | | |
#### Record Details
* Step Data (Master Step Fields)
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| --------------------------------------------- | -------------- | ----------- | ----------------------------------------- |
| Comment | Multiline Text | | |
| HML Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| RA5 Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| RNase Inhibitor Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| T4 RNA Ligase 2, Depletion Mutant Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| T4 RNA Ligase Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| 10mM ATP Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
* Step File Placeholders
* Log File - Automatically attached
* Log File - Automatically attached
* Sample Table
* Sample Display Default = Expand
* Well Sort Order = Row
* Table Columns - Global Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | LIMS ID (Container) | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Project | Project Name | Built-in | | |
### Step 2: Perform Reverse Transcription (TruSeq Small RNA v1.0)
* Master Step Name = Perform Reverse Transcription (TruSeq Small RNA v1.0.10)
* Step Type = Standard
* Derived Sample Generation = Fixed, 1
* Naming Convention = {SubmittedSampleName}
* Reagent Kits
* SuperScript II Reverse Transcriptase
* Supplier = Life Technologies
* Catalog Number = 18064-014
* Website =
* TruSeq Small RNA Library Prep Kit, Core Solutions
* Supplier = Illumina
* Catalog Number = RS-200-0012; RS-200-0024; RS-200-0036; RS-200-0048
* Website =
* TruSeq Small RNA Library Prep Kit, Indices A, B, C or D
* Supplier = Illumina
* Catalog Number = RS-200-0012; RS-200-0024; RS-200-0036; RS-200-0048
* Website =
{% hint style="info" %}
The version of Perform Reverse Transcription master step name may be different depending on the version of IPP installed.
{% endhint %}
#### Automations
Calculate Master Mix
* Trigger Location = Record Details
* Trigger Style = Automatic upon entry
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t false -h false -exp '(step.::Total samples:: = step.::Total samples:: + 1)' -log {compoundOutputFileLuid0} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t false -h false -exp '(step.::5X First Strand Buffer Volume (ul):: = step.::Total samples:: * 2.2) ; (step.::12.5 mM dNTP Mix Volume (ul):: = step.::Total samples:: * 0.55) ; (step.::100mM DTT Volume (ul):: = step.::Total samples:: * 1.1) ; (step.::RNase Inhibitor Volume (ul):: = step.::Total samples:: * 1.1) ; (step.::SuperScript II Reverse Transcriptase Volume (ul):: = step.::Total samples:: * 1.1)' -log {compoundOutputFileLuid1}"
```
{% endcode %}
Set Next Step - Advance
* Trigger Location = Record Details
* Trigger Style = Automatic upon exit
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} \
script:evaluateDynamicExpression \
-t false \
-h false \
-exp 'nextStep = ::ADVANCE::' \
-log {compoundOutputFileLuid0}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------ | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | LIMS ID (Container) | Built-in | | |
| Project | Project Name | Built-in | | |
#### Record Details
* Step Data (Master Step Fields)
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------------------------------------------------------------------------------------------- | -------------- | ----------- | ----------------------------------------- |
| Comment | Multiline Text | | |
| Dilute 25mM dNTP mix to 12.5mM (per library): 0.5ul of 25mM dNTP Mix and 0.5ul of Ultrapure water. | Toggle Switch | | Default = None Set |
| RNase Inhibitor Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| SuperScript II Reverse Transcriptase Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| 5X First Strand Buffer Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| 12.5 mM dNTP Mix Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| 100mM DTT Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
* Step File Placeholders
* Log File - Automatically attached
* Log File - Automatically attached
* Sample Table
* Sample Display Default = Expand
* Well Sort Order = Row
* Table Columns - Global Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | LIMS ID (Container) | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Project | Project Name | Built-in | | |
### Step 3: Amplify Libraries (TruSeq Small RNA v1.0)
* Master Step Name = Amplify Libraries (TruSeq Small RNA v1.0.10)
* Step Type = Add Labels
* Derived Sample Generation = Fixed, 1
* Naming Convention = {SubmittedSampleName}
* Reagent Kits
* TruSeq Small RNA Library Prep Kit, Core Solutions
* Supplier = Illumina
* Catalog Number = RS-200-0012; RS-200-0024; RS-200-0036; RS-200-0048
* Website =
* TruSeq Small RNA Library Prep Kit, Indices A, B, C or D
* Supplier = Illumina
* Catalog Number = RS-200-0012; RS-200-0024; RS-200-0036; RS-200-0048
* Website =
{% hint style="info" %}
The version of Amplify Libraries master step name may be different depending on the version of IPP installed.
{% endhint %}
#### Automations
Calculate Master Mix
* Trigger Location = Record Details
* Trigger Style = Automatic upon entry
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t false -h false -exp '(step.::Total samples:: = step.::Total samples:: + 1)' -log {compoundOutputFileLuid0} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t false -h false -exp '(step.::Ultrapure water Volume (ul):: = step.::Total samples:: * 9.35) ; (step.::PML Volume (ul):: = step.::Total samples:: * 27.5) ; (step.::RP1 Volume (ul):: = (step.::Total samples:: * 2.2).round(2)) ' -log {compoundOutputFileLuid1}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------ | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | LIMS ID (Container) | Built-in | | |
| Project | Project Name | Built-in | | |
#### Placement = Enabled
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Column
* Placement Pattern = Column
* Destination Containers
* Tube
#### Add Labels
* Label Groups
* TruSeq Small RNA
#### Record Details
* Step Data (Master Step Fields)
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------------------------------- | -------------- | ----------- | ----------------------------------------- |
| Comment | Multiline Text | | |
| Ultrapure water Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| PML Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| RP1 Volume (ul) | Numeric | | Decimal Places Displayed = 2 |
| Add 2ul of chosen RPIX to each sample. | Toggle Switch | | Default = None Set |
* Step File Placeholders
* Log File - Automatically attached
* Log File - Automatically attached
* Sample Table
* Sample Display Default = Collapse
* Well Sort Order = Row
* Table Columns - Global Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | LIMS ID (Container) | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Project | Project Name | Built-in | | |
### Step 4: Bioanalyzer QC (Library Validation) (TruSeq Small RNA v1.0)
* Master Step Name = Bioanalyzer QC (Library Validation) v2.0
* Step Type = Standard QC
* Measurement Generation = Fixed, 1
* Naming Convention = {InputItemName} Bioanalyzer
#### Automations
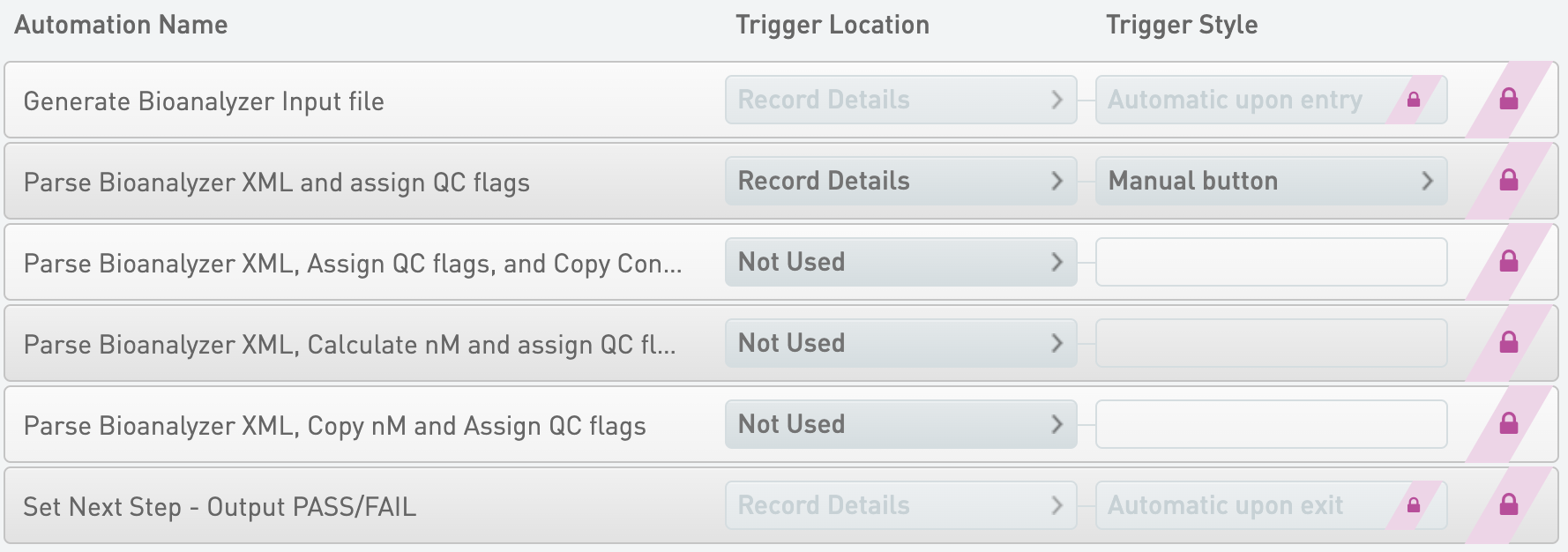
Generate Bioanalyzer Input file
* Trigger Location = Record Details
* Trigger Style = Automatic upon entry
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/DriverFileGenerator.jar script:driver_file_generator -i {processURI:v2} -u {username} -p {password} -t /opt/gls/clarity/extensions/ngs-common/v5/EPP/conf/readonly/bioA_driver_file_template.csv -o {compoundOutputFileLuid0}.csv -l {compoundOutputFileLuid1} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar script:addBlankLines -i {stepURI:v2} -u {username} -p {password} -f {compoundOutputFileLuid0}.csv -l {compoundOutputFileLuid1} -sep COMMA -b ',False,' -h 1 -c LIMSID -pre 'Sample '"
```
{% endcode %}
Parse Bioanalyzer XML and assign QC flags
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:parseBioAnalyzer -inputFile {compoundOutputFileLuid2} -log {compoundOutputFileLuid5} -configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' script:assignQC -log {compoundOutputFileLuid6} -qcResult {compoundOutputFileLuid7}"
```
{% endcode %}
Set Next Step - Output PASS/FAIL
* Trigger Location = Record Details
* Trigger Style = Automatic upon exit
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -excludeControls true -exp 'if (output.QC == true) { nextStep = ::ADVANCE:: } else { nextStep = ::ESCALATE:: }' -log {compoundOutputFileLuid0}"
```
{% endcode %}
Parse Bioanalyzer XML, Assign QC flags, and Copy Concentrations
* Trigger Location = Not Used
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:parseBioAnalyzer -inputFile {compoundOutputFileLuid2} -log {compoundOutputFileLuid5} -configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' script:assignQC -log {compoundOutputFileLuid6} -qcResult {compoundOutputFileLuid7} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'output.::Concentration:: = output.::Region 1 Conc.:: ; input.::Concentration:: = output.::Concentration:: ; output.::Conc. Units:: = ::ng/ul:: ; input.::Conc. Units:: = output.::Conc. Units::' -log {compoundOutputFileLuid8}"
```
{% endcode %}
Parse Bioanalyzer XML, Calculate nM and assign QC flags
* Trigger Location = Not Used
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:parseBioAnalyzer -inputFile {compoundOutputFileLuid2} -log {compoundOutputFileLuid5} -configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'output.::Concentration:: = output.::Region 1 Conc.:: ; output.::Molarity (nM):: = (output.::Concentration:: * 1000000) / (660 * output.::Region 1 Average Size - bp::) ; input.::Molarity (nM):: = output.::Molarity (nM):: ; output.::Conc. Units:: = ::ng/ul::' -log {compoundOutputFileLuid8} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:assignQC -log {compoundOutputFileLuid6} -qcResult {compoundOutputFileLuid7}"
```
{% endcode %}
Parse Bioanalyzer XML, Copy nM and Assign QC flags
* Trigger Location = Not Used
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:parseBioAnalyzer -inputFile {compoundOutputFileLuid2} -log {compoundOutputFileLuid5} -configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'if (output.::Conc. Units::.contains(::pg::)) {output.::Molarity (nM):: = output.::Region 1 Molarity:: / 1000} else {output.::Molarity (nM):: = output.::Region 1 Molarity::} ; (input.::Molarity (nM):: = output.::Molarity (nM)::) ' -log {compoundOutputFileLuid8} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:assignQC -log {compoundOutputFileLuid6} -qcResult {compoundOutputFileLuid7}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Row
* Sample Table (Column Headers)
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | LIMS ID (Container) | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
| Project | Project Name | Built-in | | |
#### Placement = Enabled
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Column
* Placement Pattern = Column
* Destination Containers
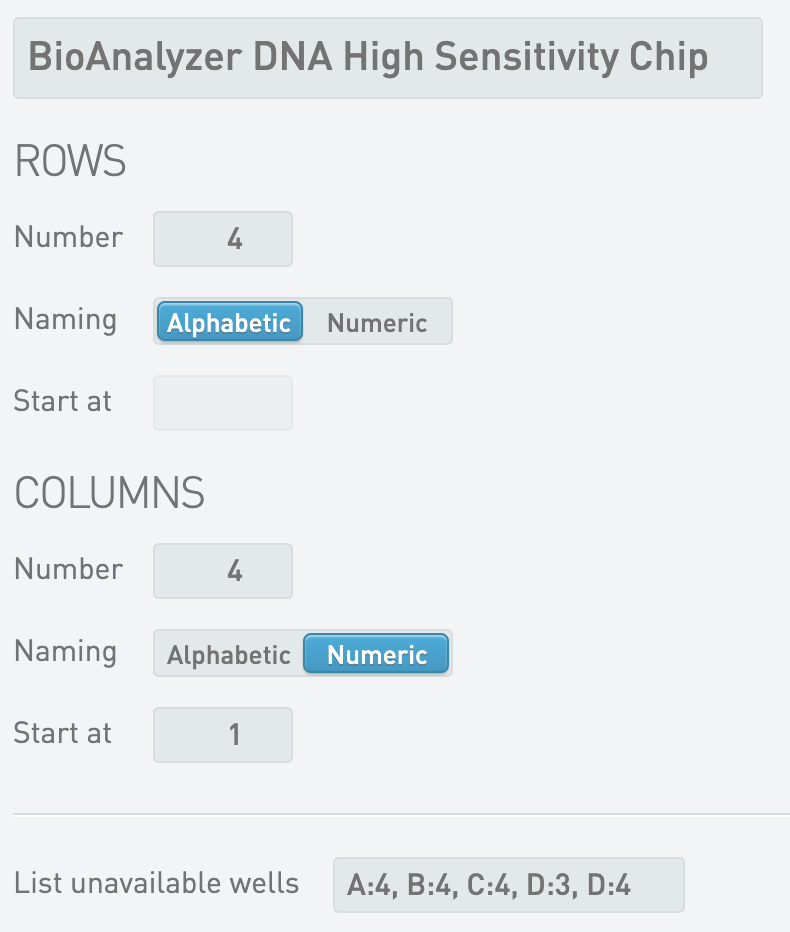
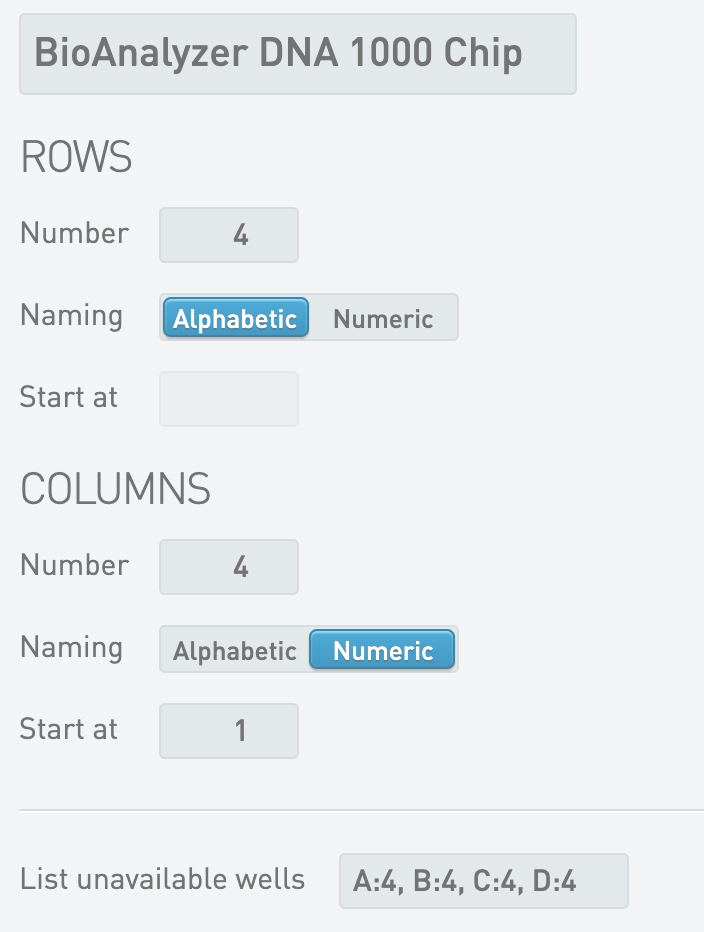
* BioAnalyzer DNA High Sensitivity Chip
Presets
|
| Criteria 1 - Source Data Field | Text Dropdown | | Presets
Concentration Conc. Units Number of Peaks found Peak 1 Size - bp Peak 1 Conc. Peak 1 Molarity Peak 2 Size - bp Peak 2 Conc. Peak 2 Molarity Peak 3 Size - bp Peak 3 Conc. Peak 3 Molarity Peak 4 Size - bp Peak 4 Conc. Peak 4 Molarity Peak 5 Size - bp Peak 5 Conc. Peak 5 Molarity Number of Regions found Region 1 Average Size - bp Region 1 Conc. Region 1 Molarity Region 2 Average Size - bp Region 2 Conc. Region 2 Molarity Region 3 Average Size - bp Region 3 Conc. Region 3 Molarity Region 4 Average Size - bp Region 4 Conc. Region 4 Molarity Region 5 Average Size - bp Region 5 Conc. Region 5 Molarity Presets
|
| Criteria 2 - Source Data Field | Text Dropdown | | Presets
Concentration Conc. Units Number of Peaks found Peak 1 Size - bp Peak 1 Conc. Peak 1 Molarity Peak 2 Size - bp Peak 2 Conc. Peak 2 Molarity Peak 3 Size - bp Peak 3 Conc. Peak 3 Molarity Peak 4 Size - bp Peak 4 Conc. Peak 4 Molarity Peak 5 Size - bp Peak 5 Conc. Peak 5 Molarity Number of Regions found Region 1 Average Size - bp Region 1 Conc. Region 1 Molarity Region 2 Average Size - bp Region 2 Conc. Region 2 Molarity Region 3 Average Size - bp Region 3 Conc. Region 3 Molarity Region 4 Average Size - bp Region 4 Conc. Region 4 Molarity Region 5 Average Size - bp Region 5 Conc. Region 5 Molarity
{% hint style="info" %}
The version of Run Gel Electrophoresis and Recover Purified Construct master step name may be different depending on the version of IPP installed.
{% endhint %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------ | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | LIMS ID (Container) | Built-in | | |
| Project | Project Name | Built-in | | |
#### Record Details
* Step Data (Master Step Fields)
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------------ | -------------- | ----------- | ----------------------------------------- |
| Comment | Multiline Text | | |
| 70% EtOH Prep Date | Date | | |
* Step File Placeholders
* Gel Image - Manually uploaded
* Sample Table
* Sample Display Default = Collapse
* Well Sort Order = Row
* Table Columns - Global Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | LIMS ID (Container) | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Project | Project Name | Built-in | | |
### Step 6: Concentrate Final Library (TruSeq Small RNA v1.0)
* Master Step Name = Concentrate Final Library (TruSeq Small RNA v1.0.10)
* Step Type = Standard
* Derived Sample Generation = Fixed, 1
* Naming Convention = {SubmittedSampleName}
* Reagent Kits
* TruSeq Small RNA Library Prep Kit, Core Solutions
* Supplier = Illumina
* Catalog Number = RS-200-0012; RS-200-0024; RS-200-0036; RS-200-0048
* Website =
{% hint style="info" %}
The version of Concentrate Final Library master step name may be different depending on the version of IPP installed.
{% endhint %}
#### Automations
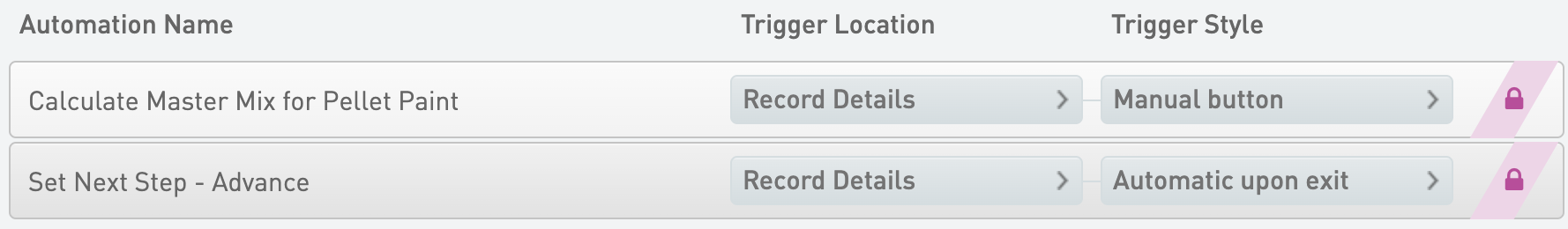
Calculate Master Mix for Pellet Paint
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp '(step.::Total samples:: = step.::Total samples:: + 1)' -log {compoundOutputFileLuid0} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'if (step.::Is Pellet Paint being used?:: == ::Yes:: ) {(step.::Optional - Ultrapure water Volume (ul):: = step.::Total samples:: * 1.98) ; (step.::Optional - 1X Pellet Paint NF Co-Precipitant Volume (ul):: = step.::Total samples:: * 0.22) ; (step.::0.1X Pellet Paint Volume (ul):: = 2)}' -log {compoundOutputFileLuid1}"
```
{% endcode %}
Set Next Step - Advance
* Trigger Location = Record Details
* Trigger Style = Automatic upon exit
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} \
script:evaluateDynamicExpression \
-t false \
-h false \
-exp 'nextStep = ::ADVANCE::' \
-log {compoundOutputFileLuid0}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------ | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | LIMS ID (Container) | Built-in | | |
| Project | Project Name | Built-in | | |
#### Record Details
* Step Data (Master Step Fields)
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------------------------------------------------- | -------------- | -------------- | ------------------------------------------------------------------- |
| Comment | Multiline Text | | |
| Glycogen Volume (ul) | Numeric | | Default = 2 Decimal Places Displayed = 0 Decimal Places Displayed = 2 Decimal Places Displayed = 2 Decimal Places Displayed = 0 Default = 30 Decimal Places Displayed = 0
Generate Bioanalyzer Input file
* Trigger Location = Record Details
* Trigger Style = Automatic upon entry
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/DriverFileGenerator.jar script:driver_file_generator -i {processURI:v2} -u {username} -p {password} -t /opt/gls/clarity/extensions/ngs-common/v5/EPP/conf/readonly/bioA_driver_file_template.csv -o {compoundOutputFileLuid0}.csv -l {compoundOutputFileLuid1} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar script:addBlankLines -i {stepURI:v2} -u {username} -p {password} -f {compoundOutputFileLuid0}.csv -l {compoundOutputFileLuid1} -sep COMMA -b ',False,' -h 1 -c LIMSID -pre 'Sample '"
```
{% endcode %}
Parse Bioanalyzer XML, Calculate nM and assign QC flags
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:parseBioAnalyzer -inputFile {compoundOutputFileLuid2} -log {compoundOutputFileLuid5} -configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'output.::Concentration:: = output.::Region 1 Conc.:: ; output.::Molarity (nM):: = (output.::Concentration:: * 1000000) / (660 * output.::Region 1 Average Size - bp::) ; input.::Molarity (nM):: = output.::Molarity (nM):: ; output.::Conc. Units:: = ::ng/ul::' -log {compoundOutputFileLuid8} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:assignQC -log {compoundOutputFileLuid6} -qcResult {compoundOutputFileLuid7}"
```
{% endcode %}
Set Next Step - Output PASS/FAIL
* Trigger Location = Record Details
* Trigger Style = Automatic upon exit
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -excludeControls true -exp 'if (output.QC == true) { nextStep = ::ADVANCE:: } else { nextStep = ::ESCALATE:: }' -log {compoundOutputFileLuid0}"
```
{% endcode %}
Parse Bioanalyzer XML and assign QC flags
* Trigger Location = Not Used
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:parseBioAnalyzer -inputFile {compoundOutputFileLuid2} -log {compoundOutputFileLuid5} -configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' script:assignQC -log {compoundOutputFileLuid6} -qcResult {compoundOutputFileLuid7}"
```
{% endcode %}
Parse Bioanalyzer XML, Assign QC flags, and Copy Concentrations
* Trigger Location = Not Used
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:parseBioAnalyzer -inputFile {compoundOutputFileLuid2} -log {compoundOutputFileLuid5} -configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' script:assignQC -log {compoundOutputFileLuid6} -qcResult {compoundOutputFileLuid7} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'output.::Concentration:: = output.::Region 1 Conc.:: ; input.::Concentration:: = output.::Concentration:: ; output.::Conc. Units:: = ::ng/ul:: ; input.::Conc. Units:: = output.::Conc. Units::' -log {compoundOutputFileLuid8}"
```
{% endcode %}
Parse Bioanalyzer XML, Copy nM and Assign QC flags
* Trigger Location = Not Used
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:parseBioAnalyzer -inputFile {compoundOutputFileLuid2} -log {compoundOutputFileLuid5} -configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'if (output.::Conc. Units::.contains(::pg::)) {output.::Molarity (nM):: = output.::Region 1 Molarity:: / 1000} else {output.::Molarity (nM):: = output.::Region 1 Molarity::} ; (input.::Molarity (nM):: = output.::Molarity (nM)::) ' -log {compoundOutputFileLuid8} && /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {processURI:v2} -u {username} -p {password} script:assignQC -log {compoundOutputFileLuid6} -qcResult {compoundOutputFileLuid7}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Row
* Sample Table (Column Headers)
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | LIMS ID (Container) | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
| Project | Project Name | Built-in | | |
#### Placement = Enabled
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Column
* Placement Pattern = Column
* Destination Containers
* BioAnalyzer DNA High Sensitivity Chip
Nextera DNA Flex Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 150.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 1,500.00
Nextera Mate Pair Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Average Size - bp
* Criteria 1 - Threshold Value = 150.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Average Size - bp
* Criteria 2 - Threshold Value = 400.00
Nextera XT DNA Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 250.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 1,000.00
NRCC Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 350.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 1,000.00
TruSeq ChIP-Seq Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Average Size - bp
* Criteria 1 - Threshold Value = 150.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Average Size - bp
* Criteria 2 - Threshold Value = 400.00
TruSeq Exome Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 150.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 1,000.00
TruSeq Methyl Capture EPIC Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 200.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 300.00
TruSeq Rapid Exome Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 200.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 500.00
TruSeq RNA Access Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 200.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 320.00
TruSeq RNA Exome Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 200.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 320.00
TruSeq Small RNA Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Average Size - bp
* Criteria 1 - Threshold Value = 100.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Average Size - bp
* Criteria 2 - Threshold Value = 200.00
TruSeq Stranded mRNA Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Average Size - bp
* Criteria 1 - Threshold Value = 250.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Average Size - bp
* Criteria 2 - Threshold Value = 275.00
TruSeq Stranded Total RNA Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Average Size - bp
* Criteria 1 - Threshold Value = 250.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Average Size - bp
* Criteria 2 - Threshold Value = 275.00
TruSeq Targeted RNA Expression Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 100.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 300.00
TruSight Myeloid Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Average Size - bp
* Criteria 1 - Threshold Value = 150.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Size - bp
* Criteria 2 - Threshold Value = 400.00
TruSight RNA Fusion Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Average Size - bp
* Criteria 1 - Threshold Value = 160.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Size - bp
* Criteria 2 - Threshold Value = 700.00
TSCA Library Validation
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Average Size - bp
* Criteria 1 - Threshold Value = 300.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Size - bp
* Criteria 2 - Threshold Value = 400.00
* Step Data
* Group of Defaults = TruSeq Small RNA Library Validation
* Master Step Fields
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------------------------------------- | -------------- | -------------- | --------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
| Criteria 1 - Operator | Text Dropdown | Custom Entries | Presets
|
| Criteria 1 - Source Data Field | Text Dropdown | | Presets
Concentration Conc. Units Number of Peaks found Peak 1 Size - bp Peak 1 Conc. Peak 1 Molarity Peak 2 Size - bp Peak 2 Conc. Peak 2 Molarity Peak 3 Size - bp Peak 3 Conc. Peak 3 Molarity Peak 4 Size - bp Peak 4 Conc. Peak 4 Molarity Peak 5 Size - bp Peak 5 Conc. Peak 5 Molarity Number of Regions found Region 1 Average Size - bp Region 1 Conc. Region 1 Molarity Region 2 Average Size - bp Region 2 Conc. Region 2 Molarity Region 3 Average Size - bp Region 3 Conc. Region 3 Molarity Region 4 Average Size - bp Region 4 Conc. Region 4 Molarity Region 5 Average Size - bp Region 5 Conc. Region 5 Molarity Presets
|
| Criteria 2 - Source Data Field | Text Dropdown | | Presets
Concentration Conc. Units Number of Peaks found Peak 1 Size - bp Peak 1 Conc. Peak 1 Molarity Peak 2 Size - bp Peak 2 Conc. Peak 2 Molarity Peak 3 Size - bp Peak 3 Conc. Peak 3 Molarity Peak 4 Size - bp Peak 4 Conc. Peak 4 Molarity Peak 5 Size - bp Peak 5 Conc. Peak 5 Molarity Number of Regions found Region 1 Average Size - bp Region 1 Conc. Region 1 Molarity Region 2 Average Size - bp Region 2 Conc. Region 2 Molarity Region 3 Average Size - bp Region 3 Conc. Region 3 Molarity Region 4 Average Size - bp Region 4 Conc. Region 4 Molarity Region 5 Average Size - bp Region 5 Conc. Region 5 Molarity
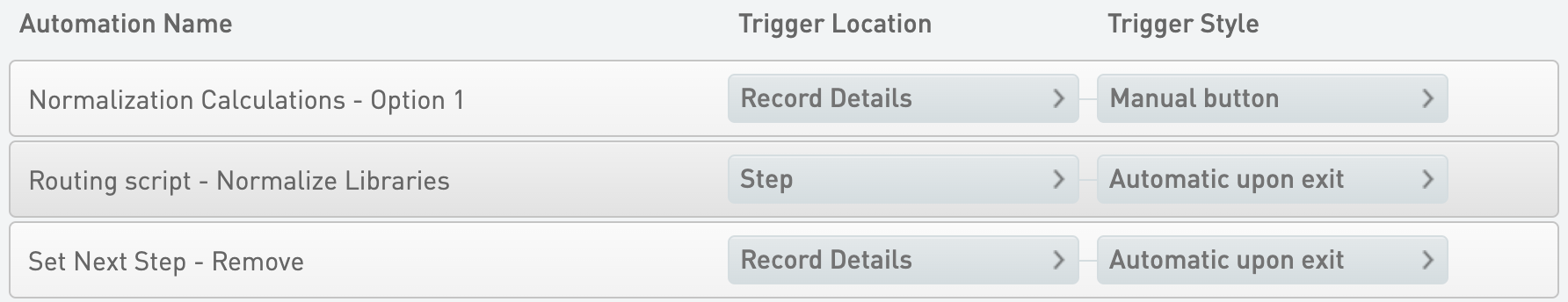
Normalization Calculations - Option 1
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'output.::Molarity (nM):: = input.::Molarity (nM):: ; if (output.::Molarity (nM):: <= step.::Target Normalization (nM)::) {output.::Sample Volume (ul):: = step.::Final Volume (ul):: ; output.::Buffer Volume (ul):: = 0 ; output.::Normalized Molarity (nM):: = output.::Molarity (nM)::} else {output.::Sample Volume (ul):: = (step.::Target Normalization (nM):: * step.::Final Volume (ul):: ) / input.::Molarity (nM):: ; output.::Buffer Volume (ul):: = step.::Final Volume (ul):: - output.::Sample Volume (ul):: ; output.::Normalized Molarity (nM):: = step.::Target Normalization (nM)::}' -log {compoundOutputFileLuid0}"
```
{% endcode %}
Set Next Step - Remove
* Trigger Location = Record Details
* Trigger Style = Automatic upon exit
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t false -h false -exp 'nextStep = ::REMOVE::' -log {compoundOutputFileLuid0}"
```
{% endcode %}
Routing script - Normalize Libraries
* Trigger Location = Step
* Trigger Style = Automatic upon exit
{% code overflow="wrap" %}
```markup
bash -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -u {username} -p {password} -i {stepURI:v2} -l {compoundOutputFileLuid0} script:changeWorkflow \
\
--FIELD_NAME 'Sequencing Instrument' \
--FIELD_VALUE 'MiSeq' \
--WORKFLOW 'MiSeq Sequencing v3.2' \
--STEP 'Library Pooling (MiSeq v3.2)' \
--INPUTS_OR_OUTPUTS 'OUTPUTS' \
\
--FIELD_NAME 'Sequencing Instrument' \
--FIELD_VALUE 'NextSeq' \
--WORKFLOW 'NextSeq 500/550 Sequencing v1.2' \
--STEP 'Library Pooling (NextSeq 500/550 v1.2)' \
--INPUTS_OR_OUTPUTS 'OUTPUTS' \
\
--FIELD_NAME 'Sequencing Instrument' \
--FIELD_VALUE 'NovaSeq 2.0' \
--WORKFLOW 'NovaSeq 6000 v2.3' \
--STEP 'Define Run Format (NovaSeq 6000 v2.3)' \
--INPUTS_OR_OUTPUTS 'OUTPUTS' \
\
--FIELD_NAME 'Sequencing Instrument' \
--FIELD_VALUE 'NovaSeq 3.0' \
--WORKFLOW 'NovaSeq 6000 v3.8' \
--STEP 'Define Run Format (NovaSeq 6000 v3.8)' \
--INPUTS_OR_OUTPUTS 'OUTPUTS' \
\
--FIELD_NAME 'Sequencing Instrument' \
--FIELD_VALUE 'NovaSeqDx' \
--WORKFLOW 'NovaSeqDx v1.2' \
--STEP 'Define Run Format (NovaSeqDx v1.2)' \
--INPUTS_OR_OUTPUTS 'OUTPUTS' \
\
--FIELD_NAME 'Sequencing Instrument' \
--FIELD_VALUE 'NextSeq 1000/2000' \
--WORKFLOW 'NextSeq 1000/2000 Sequencing v2.4' \
--STEP 'Library Pooling and Dilution (NextSeq 1000/2000 Sequencing v2.4)' \
--INPUTS_OR_OUTPUTS 'OUTPUTS' \
\
--FIELD_NAME 'Sequencing Instrument' \
--FIELD_VALUE 'NovaSeq X Series' \
--WORKFLOW 'NovaSeq X Series v1.1' \
--STEP 'Assign Analysis Configuration Template (NovaSeq X Series Sequencing v1.1)' \
--INPUTS_OR_OUTPUTS 'OUTPUTS' \
\
--FIELD_NAME 'Sequencing Instrument' \
--FIELD_VALUE 'NextSeq 1000/2000 On-Prem' \
--WORKFLOW 'NextSeq 1000/2000 On-Prem Sequencing v1.0' \
--STEP 'Library Pooling and Dilution (NextSeq 1000/2000 On-Prem Sequencing v1.0)' \
--INPUTS_OR_OUTPUTS 'OUTPUTS'"
```
{% endcode %}
> ℹ The field value and actual version of the workflows and steps in the routing automation script may be different depending on the version of IPP installed.
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Containers
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------ | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | LIMS ID (Container) | Built-in | | |
| Project | Project Name | Built-in | | |
#### Record Details
* Step Data (Master Step Fields)
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------------------- | -------------- | -------------- | ------------------------------------------------------------------ |
| Comment | Multiline Text | | |
| Final Volume (ul) | Numeric | Required Field | Decimal Places Displayed = 2 Default = 2 Decimal Places Displayed = 2 Presets
MiSeq NextSeq NextSeq 1000/2000 NextSeq 1000/2000 On-Prem NovaSeq 2.0 NovaSeq 3.0 NovaSeq X Series NovaSeqDx
```
The question should be specific, self-contained, and written in natural language.
The response will contain a direct answer to the question and relevant excerpts and sources from the documentation.
Use this mechanism when the answer is not explicitly present in the current page, you need clarification or additional context, or you want to retrieve related documentation sections.