
Figure 1. Accessing Chromosome view via the context-sensitive menu (the content of the Visualisation section depends on the selected data node)

Figure 1. Accessing Chromosome view via the context-sensitive menu (the content of the Visualisation section depends on the selected data node)
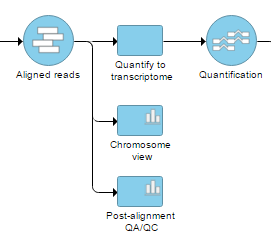
Figure 2. Selecting Chromosome view from the context-sensitive menu adds a Chromosome view task node to the canvas. To open the view, on it
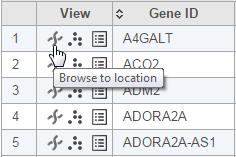
Figure 3. Accessing the Chromosome view from results table (mouse-over balloon is visible when hovering over the chromosome icon). The image is an example, based on gene expression pipeline