# Library Validation QC 5.1.2
## Overview
The Library Validation QC protocol contains steps to track quality control data from commonly used QC instruments. Through data integrations, QC data can be added to Clarity LIMS directly from the instrument outputs.
Highlights of this protocol include:
* Data integrations for Bioanalyzer 2100 and Tapestation 2200, including creation of the input file and parsing of data output.
* Possible integrations with qPCR instruments: TaqMan 7900 or ViiA7.
* Assignment of pass/fail QC based on chosen criteria.
* Aggregation of data from multiple QC steps.
## Protocol 1: Library Validation QC 5.1.2
* Protocol Type = QC
* Capacity = 50
* QC Filters
* Bioanalyzer QC (DNA) 5.1.2
* Did not pass - Bioanalyzer QC (DNA) 5.1.2
* qPCR QC 5.1.2
* Did not pass - qPCR QC 5.1.2
* Tapestation QC (DNA) 5.1.2
* Did not pass - Tapestation QC (DNA) 5.1.2
* Aggregate QC (Library Validation) 5.1.2
* Passed - Bioanalyzer QC (DNA) 5.1.2 OR
* Passed - qPCR QC 5.1.2 OR
* Passed - Tapestation QC (DNA) 5.1.2
### Step: Bioanalyzer QC (DNA) 5.1.2
* Master Step Name = Bioanalyzer QC (DNA) 5.1.2
* Step Type = Standard QC
* Measurement Generation = Fixed, 1
* Naming Convention = {InputItemName} Bioanalyzer
#### Automations
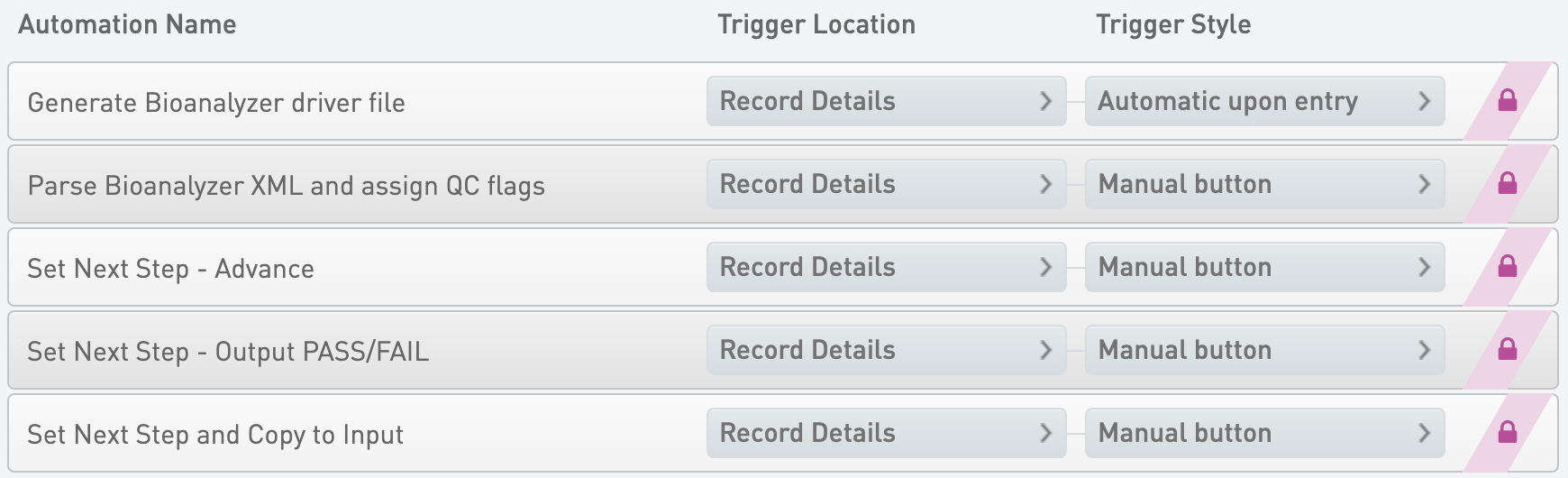
Generate Bioanalyzer driver file
* Trigger Location = Record Details
* Trigger Style = Automatic upon entry
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/DriverFileGenerator.jar -u {username} -p {password} \
script:driver_file_generator \
-i {processURI:v2} \
-t /opt/gls/clarity/extensions/ngs-common/v5/EPP/conf/readonly/bioA_driver_file_template.csv \
-o {compoundOutputFileLuid0}.csv \
-l {compoundOutputFileLuid1} \
&& /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -u {username} -p {password} \
script:addBlankLines \
-i {stepURI:v2} \
-f {compoundOutputFileLuid0}.csv \
-l {compoundOutputFileLuid1} \
-sep COMMA \
-b ',False,' \
-h 1 \
-c LIMSID \
-pre 'Sample '"
```
{% endcode %}
Parse Bioanalyzer XML and assign QC flags
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -u {username} -p {password} -i {processURI:v2} \
script:parseBioAnalyzer \
-inputFile {compoundOutputFileLuid2} \
-log {compoundOutputFileLuid5} \
-configFile '/opt/gls/clarity/extensions/conf/v5/bioanalyzer/defaultBioAnalyzerDNAConfig.groovy' \
script:assignQC \
-log {compoundOutputFileLuid6} \
-qcResult {compoundOutputFileLuid7}"
```
{% endcode %}
Set Next Step - Advance
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} \
script:evaluateDynamicExpression \
-t true \
-h false \
-exp 'nextStep = ::ADVANCE::' \
-log {compoundOutputFileLuid0}"
```
{% endcode %}
Set Next Step - Output PASS/FAIL
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -excludeControls true -exp 'if (output.QC == true) { nextStep = ::ADVANCE:: } else { nextStep = ::ESCALATE:: }' -log {compoundOutputFileLuid0}"
```
{% endcode %}
Set Next Step and Copy to Input
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -i {stepURI:v2} -u {username} -p {password} script:evaluateDynamicExpression -t true -h false -exp 'nextStep = ::ADVANCE:: ; input.::Concentration (ng/ul):: = output.::Concentration::' -log {compoundOutputFileLuid7}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Previous Steps
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ---------------- | ------------------- | -------------- | -------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
| Derived Sample | Workflow | Text | | |
| Project | Project Name | Built-in | | |
| Submitted Sample | Application | Text Dropdown | Custom Entries | Presets
TruSeq mRNA sequencing TruSeq DNA sequencing (large genome de novo) TruSeq DNA sequencing (large genome re-seq) TruSeq DNA sequencing (small genome de novo) TruSeq DNA sequencing (small genome re-seq) Nextera DNA sequencing TruSeq Custom Amplicon sequencing ChIP-sequencing Exome sequencing Mate pair sequencing Small RNA Sequencing Presets
|
| Submitted Sample | Read Length | Text | | |
| Submitted Sample | Sample Type | Text | | |
| Submitted Sample | Sequencing Coverage | Text | | |
| Submitted Sample | Sequencing Method | Text Dropdown | Custom Entries | Presets
Single Read Paired End Read Indexed Single Read Indexed Paired End Read
NRCC
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 150.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 1,000.00
* Use strict matching for Bioanalyzer results = No
Peak 2 Size Thresholds
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 100.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 1,000.00
* Use strict matching for Bioanalyzer results = No
TruSeq ChIP-Seq
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Region 1 Molarity
* Criteria 1 - Threshold Value = 5.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Region 1 Molarity
* Criteria 2 - Threshold Value = 10.00
* Use strict matching for Bioanalyzer results = No
TruSeq Methyl Capture EPIC
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 100.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 300.00
TruSeq Rapid Exome
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 Size - bp
* Criteria 1 - Threshold Value = 150.00
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 Size - bp
* Criteria 2 - Threshold Value = 1,000.00
* Use strict matching for Bioanalyzer results = No
* Step Data
* Step Data Heading = Begin by uploading the required Bioanalyzer XML result file
* Group of Defaults = Peak 2 Size Thresholds
* Master Step Fields
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------------------------------------- | -------------- | ----------- | --------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
| Criteria 1 - Operator | Text Dropdown | | Presets
|
| Criteria 1 - Source Data Field | Text Dropdown | | Presets
Concentration Conc. Units Number of Peaks found Peak 1 Size - bp Peak 1 Conc. Peak 1 Molarity Peak 2 Size - bp Peak 2 Conc. Peak 2 Molarity Peak 3 Size - bp Peak 3 Conc. Peak 3 Molarity Peak 4 Size - bp Peak 4 Conc. Peak 4 Molarity Peak 5 Size - bp Peak 5 Conc. Peak 5 Molarity Number of Regions found Region 1 Average Size - bp Region 1 Conc. Region 1 Molarity Region 2 Average Size - bp Region 2 Conc. Region 2 Molarity Region 3 Average Size - bp Region 3 Conc. Region 3 Molarity Region 4 Average Size - bp Region 4 Conc. Region 4 Molarity Region 5 Average Size - bp Region 5 Conc. Region 5 Molarity Presets
|
| Criteria 2 - Source Data Field | Text Dropdown | | Presets
Concentration Conc. Units Number of Peaks found Peak 1 Size - bp Peak 1 Conc. Peak 1 Molarity Peak 2 Size - bp Peak 2 Conc. Peak 2 Molarity Peak 3 Size - bp Peak 3 Conc. Peak 3 Molarity Peak 4 Size - bp Peak 4 Conc. Peak 4 Molarity Peak 5 Size - bp Peak 5 Conc. Peak 5 Molarity Number of Regions found Region 1 Average Size - bp Region 1 Conc. Region 1 Molarity Region 2 Average Size - bp Region 2 Conc. Region 2 Molarity Region 3 Average Size - bp Region 3 Conc. Region 3 Molarity Region 4 Average Size - bp Region 4 Conc. Region 4 Molarity Region 5 Average Size - bp Region 5 Conc. Region 5 Molarity
Assign QC flags
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -u {username} -p {password} \
script:assignQC \
-i {processURI:v2} \
-log {compoundOutputFileLuid1} \
-qcResult {compoundOutputFileLuid2}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Previous Steps
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ---------------- | ------------------- | -------------- | -------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
| Derived Sample | Workflow | Text | | |
| Project | Project Name | Built-in | | |
| Submitted Sample | Application | Text Dropdown | Custom Entries | Presets
TruSeq mRNA sequencing TruSeq DNA sequencing (large genome de novo) TruSeq DNA sequencing (large genome re-seq) TruSeq DNA sequencing (small genome de novo) TruSeq DNA sequencing (small genome re-seq) Nextera DNA sequencing TruSeq Custom Amplicon sequencing ChIP-sequencing Exome sequencing Mate pair sequencing Small RNA Sequencing Presets
|
| Submitted Sample | Read Length | Text | | |
| Submitted Sample | Sample Type | Text | | |
| Submitted Sample | Sequencing Coverage | Text | | |
| Submitted Sample | Sequencing Method | Text Dropdown | Custom Entries | Presets
Single Read Paired End Read Indexed Single Read Indexed Paired End Read
Concentration Thresholds
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Concentration
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Concentration
* Step Data
* Step Data Heading = Begin by completing the Sample Details table
* Group of Defaults = Concentration Thresholds
* Master Step Fields
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------------------------ | -------------- | ----------- | ------------------------------------------------------------------ |
| Criteria 1 - Operator | Text Dropdown | | Presets
|
| Criteria 1 - Source Data Field | Text Dropdown | | Presets
|
| Criteria 1 - Threshold Value | Numeric | | Decimal Places Displayed = 2 |
| Criteria 2 - Operator | Text Dropdown | | Presets
|
| Criteria 2 - Source Data Field | Text Dropdown | | Presets
|
| Criteria 2 - Threshold Value | Numeric | | Decimal Places Displayed = 2 |
* Step File Placeholders
* qPCR Result File - Manually uploaded
* QC Assignment Log File - Automatically attached
* QC Assignment Report - Automatically attached
* Sample Table
* Enable QC Flags = Yes
* Sample Display Default = Collapse
* Well Sort Order = Row
* File Column Options
* File Column Display = Hide
* File Attachment Method = Auto
* Table Columns - Global Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Measurement | Concentration | Numeric | | Decimal Places Displayed = 2 |
| Measurement | Conc. Units | Text | | |
| Project | Project Name | Built-in | | |
### Step: Tapestation QC (DNA) 5.1.2
* Master Step Name = Tapestation QC (DNA) 5.1.2
* Step Type = Standard QC
* Measurement Generation = Fixed, 1
* Naming Convention = {InputItemName} Tapestation
#### Automations
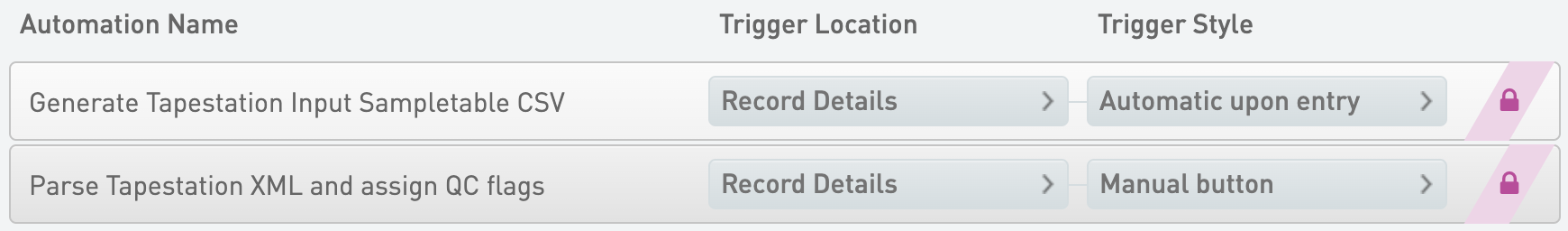
Generate Tapestation Input Sampletable CSV
* Trigger Location = Record Details
* Trigger Style = Automatic upon entry
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/DriverFileGenerator.jar -u {username} -p {password} \
script:driver_file_generator \
-i {processURI:v2} \
-t /opt/gls/clarity/extensions/ngs-common/v5/EPP/conf/readonly/tapestation_driver_file_template.csv \
-o {compoundOutputFileLuid0}.csv \
-l {compoundOutputFileLuid1}"
```
{% endcode %}
Parse Tapestation XML and assign QC flags
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/XmlSampleNameParser.jar -u {username} -p {password} \
script:parseXmlBySampleName \
-i {processURI:v2} \
-inputFile {compoundOutputFileLuid2} \
-log {compoundOutputFileLuid3}.html \
-configFile /opt/gls/clarity/extensions/conf/v5/tapestation/defaultTapeStationDNAConfig.groovy \
&& /opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -u {username} -p {password} \
script:assignQC \
-i {processURI:v2} \
-log {compoundOutputFileLuid4} \
-qcResult {compoundOutputFileLuid5}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Previous Steps
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Container Name | Built-in | | |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------ | ------------------- | -------------- | ----------- | ----------------------------------------- |
| Container | LIMS ID (Container) | Built-in | | |
| Project | Project Name | Built-in | | |
#### Record Details
**Group of Defaults**
Peak 2 Size Thresholds
* Criteria 1 - Operator = >=
* Criteria 1 - Source Data Field = Peak 2 MW
* Criteria 1 - Threshold Value = 100
* Criteria 2 - Operator = <=
* Criteria 2 - Source Data Field = Peak 2 MW
* Criteria 2 - Threshold Value = 1,000
* Step Data
* Group of Defaults = Peak 2 Size Thresholds
* Master Step Fields
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ------------------------------ | -------------- | ----------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
| Criteria 1 - Operator | Text Dropdown | | |
| Criteria 1 - Source Data Field | Text Dropdown | | Presets
Concentration Conc. Units Molarity Units MW Units Peak 1 % Integrated Area Peak 1 Conc. Peak 1 MW Peak 2 % Integrated Area Peak 2 Conc. Peak 2 MW Peak 3 % Integrated Area Peak 3 Conc. Peak 3 MW Peak 4 % Integrated Area Peak 4 Conc. Peak 4 MW Peak 5 % Integrated Area Peak 5 Conc. Peak 5 MW Region 1 % of Total Region 1 Average Size - bp Region 1 Conc. Region 1 Molarity Region 2 % of Total Region 2 Average Size - bp Region 2 Conc. Region 2 Molarity Region 3 % of Total Region 3 Average Size - bp Region 3 Conc. Region 3 Molarity Region 4 % of Total Region 4 Average Size - bp Region 4 Conc. Region 4 Molarity Region 5 % of Total Region 5 Average Size - bp Region 5 Conc. Region 5 Molarity Default = Concentration Decimal Places Displayed = 0 Presets
Conc. Units Peak 1 Conc. Peak 2 Conc. Peak 3 Conc. Peak 4 Conc. Peak 5 Conc. Region 1 Average Size - bp Region 1 Conc. Region 1 Molarity Region 2 Average Size - bp Region 2 Conc. Region 2 Molarity Region 3 Average Size - bp Region 3 Conc. Region 3 Molarity Region 4 Average Size - bp Region 4 Conc. Region 4 Molarity Region 5 Average Size - bp Region 5 Conc. Region 5 Molarity Region 1 % of Total Region 2 % of Total Region 3 % of Total Region 4 % of Total Region 5 % of Total Peak 1 MW Peak 2 MW Peak 3 MW Peak 4 MW Peak 5 MW Molarity Units Peak 1 % Integrated Area Peak 2 % Integrated Area Peak 3 % Integrated Area Peak 4 % Integrated Area Peak 5 % Integrated Area MW Units Concentration Default = Conc. Units Decimal Places Displayed = 0
Aggregate QC Flags and Copy Fields
* Trigger Location = Record Details
* Trigger Style = Manual button
{% code overflow="wrap" %}
```markup
bash -l -c "/opt/gls/clarity/bin/java -jar /opt/gls/clarity/extensions/ngs-common/v5/EPP/ngs-extensions.jar -u {username} -p {password} \
script:setUDF \
-i {processURI:v2} \
-f 'Progress' \
-t '//input/@uri->//sample/@uri' \
-v 'Library preparation and QC validation complete' \
script:aggregateQC \
-i {stepURI:v2} \
-c 'true' \
-a 'true' \
-log {compoundOutputFileLuid0} \
-aggregatelog {compoundOutputFileLuid1} \
-copylog {compoundOutputFileLuid2}"
```
{% endcode %}
#### Queue/Ice Bucket
* Defaults
* Sample Grouping = Group by Previous Steps
* Well Sort Order = Row
* Sample Table
* Column Headers
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------- | -------------- | -------------- | ----------- | ----------------------------------------- |
| Container | Well | Built-in | | |
| Derived Sample | Sample Name | Built-in | | |
| Derived Sample | Waiting | Built-in | | |
| Derived Sample | Workflow | Text | | |
| Project | Project Name | Built-in | | |
* Expanded View Fields
| **Category** | **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| ---------------- | ------------------- | -------------- | -------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
| Container | Container Name | Built-in | | |
| Submitted Sample | Application | Text Dropdown | Custom Entries | Presets
TruSeq mRNA sequencing TruSeq DNA sequencing (large genome de novo) TruSeq DNA sequencing (large genome re-seq) TruSeq DNA sequencing (small genome de novo) TruSeq DNA sequencing (small genome re-seq) Nextera DNA sequencing TruSeq Custom Amplicon sequencing ChIP-sequencing Exome sequencing Mate pair sequencing Small RNA Sequencing Presets
|
| Submitted Sample | Read Length | Text | | |
| Submitted Sample | Sample Type | Text | | |
| Submitted Sample | Sequencing Coverage | Text | | |
| Submitted Sample | Sequencing Method | Text Dropdown | Custom Entries | Presets
Single Read Paired End Read Indexed Single Read Indexed Paired End Read
Tapestation Values
* Bioanalzyer QC (DNA) 5.1.2 = Use if available (Priority 5)
* Copy task 1 - Source Field = Concentration
* Copy task 1 - Source Step = Tapestation QC (DNA) 5.1.2
* Copy task 2 - Source Field = Conc. Units
* Copy task 2 - Source Step = Tapestation QC (DNA) 5.1.2
* NanoDrop QC (DNA) 5.1.2 = Use if available (Priority 5)
* PicoGreen QC (DNA) 5.1.2 = Use if available (Priority 5)
* qPCR QC 5.1.2 = Use if available (Priority 5)
* Qubit QC (DNA) 5.1.2 = Use if available (Priority 5)
* Tapestation QC (DNA) 5.1.2 = Use if available (Priority 5)
* Step Data
* Step Data Heading = Begin by verifying copy tasks and aggregation priorities
* Group of Defaults = Tapestation Values
* Master Step Fields
| **Field Name** | **Field Type** | **Options** | **Additional Options and Dropdown Items** |
| -------------------------- | -------------- | -------------- | ------------------------------------------------------------------------------------------------------------------- |
| Copy task 1 - Source Field | Text Dropdown | | Presets
Concentration Conc. Units A260/280 ratio Presets
Bioanalyzer QC (DNA) 5.1.2 qPCR QC 5.1.2 Tapestation QC (DNA) 5.1.2 Presets
Concentration Conc. Units A260/280 ratio Presets
Bioanalyzer QC (DNA) 5.1.2 qPCR QC 5.1.2 Tapestation QC (DNA) 5.1.2 Presets
Concentration Conc. Units A260/280 ratio Presets
Bioanalyzer QC (DNA) 5.1.2 qPCR QC 5.1.2 Tapestation QC (DNA) 5.1.2
```
The question should be specific, self-contained, and written in natural language.
The response will contain a direct answer to the question and relevant excerpts and sources from the documentation.
Use this mechanism when the answer is not explicitly present in the current page, you need clarification or additional context, or you want to retrieve related documentation sections.